Permutation Results on LUHMES CROP-seq Data
– In-house scripts
Yifan Zhou (zhouyf@uchicago.edu)
2022-09-19
Here, we summarize the permutation results for GSFA, scMAGeCK, DESeq2, and MAST.
1 GSFA Permutation
Cell labels in the expression data were permuted randomly so that they are no longer correlated with the knock-down conditions. Then GSFA was performed still using all conditions as guides. Factor-guide association as well as the LFSR of each gene were evaluated as usual.
In total, 10 random permutation rounds like this were conducted. In each case, Gibbs sampling was conducted for 3000 iterations, and the posterior mean estimates were averaged over the last 1000 iterations. SVD Initialization.
1.1 Factor ~ KO Association
1.1.1 Posterior Mean of Beta
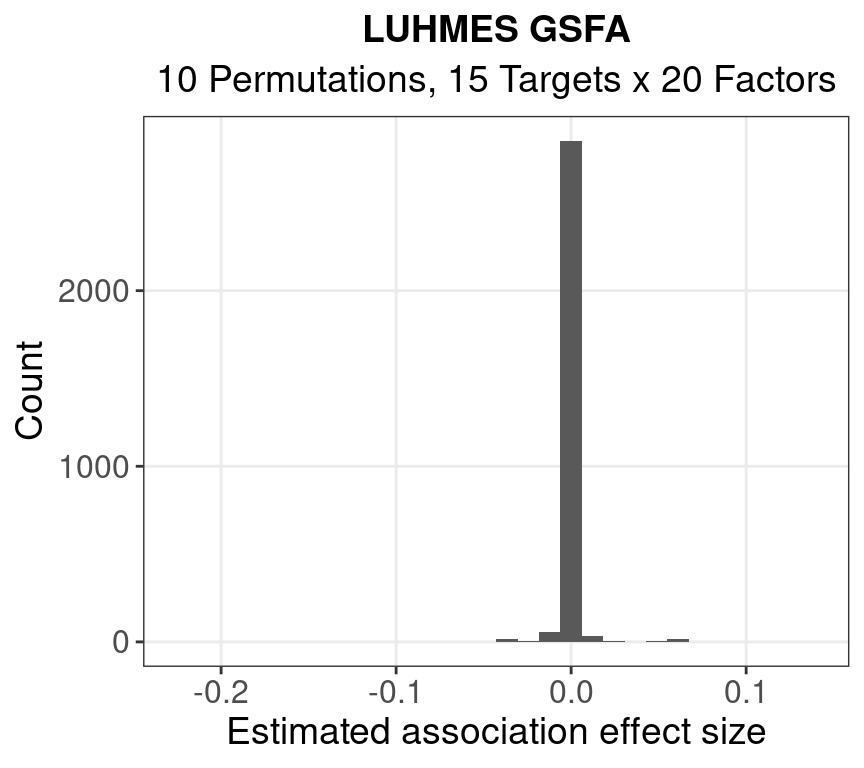 #
of |beta|’s > 0.05: 26
#
of |beta|’s > 0.05: 26
1.1.2 Beta PIP
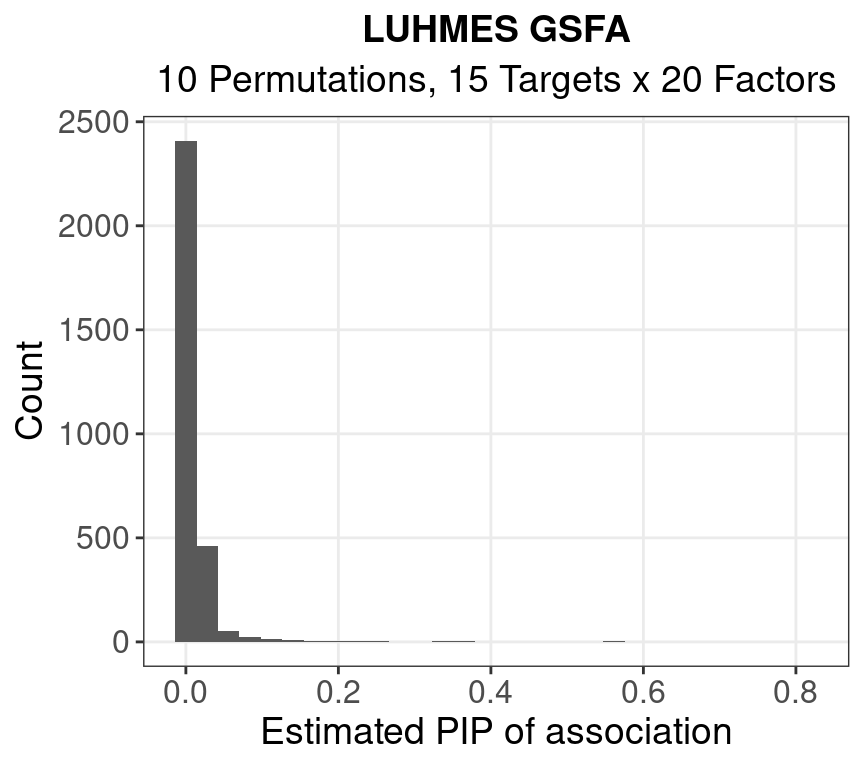 #
of PIPs > 0.9: 0
#
of PIPs > 0.9: 0
1.2 LFSR
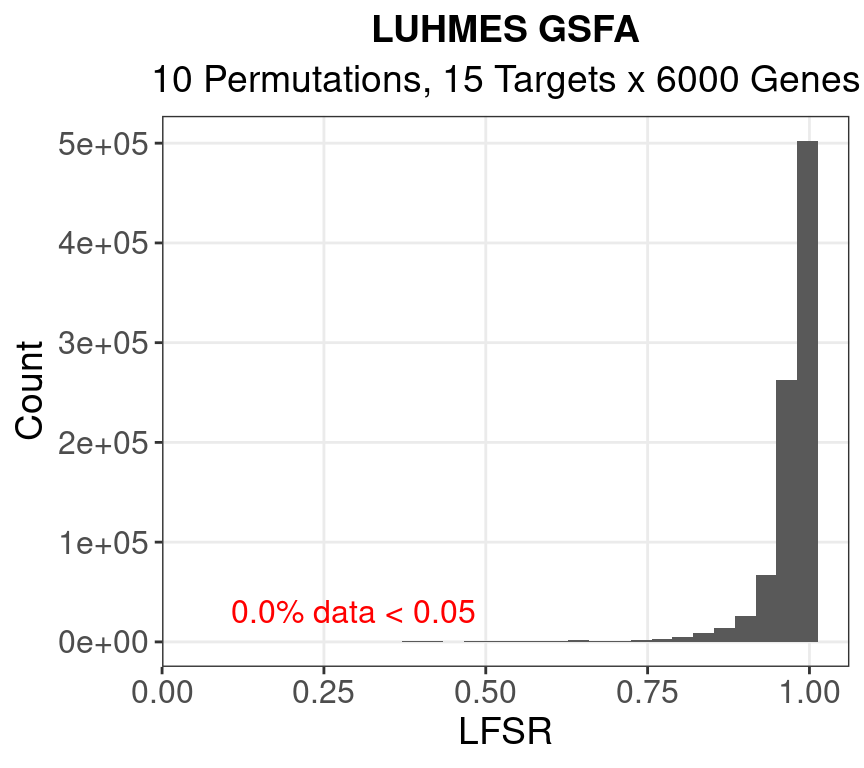
2 scMAGeCK Permutation
Cell labels in the expression data were permuted randomly so that they are no longer correlated with the knock-down conditions. Then scMAGeCK-LR test was performed for all guides at once.
In total, 10 random permutation rounds like this were conducted.
The outputs are empirical p values, some of them equal to 0 exactly, and had to be replaced with 0.0005 for QQ-plot.
2.1 Combined from 10 permutations
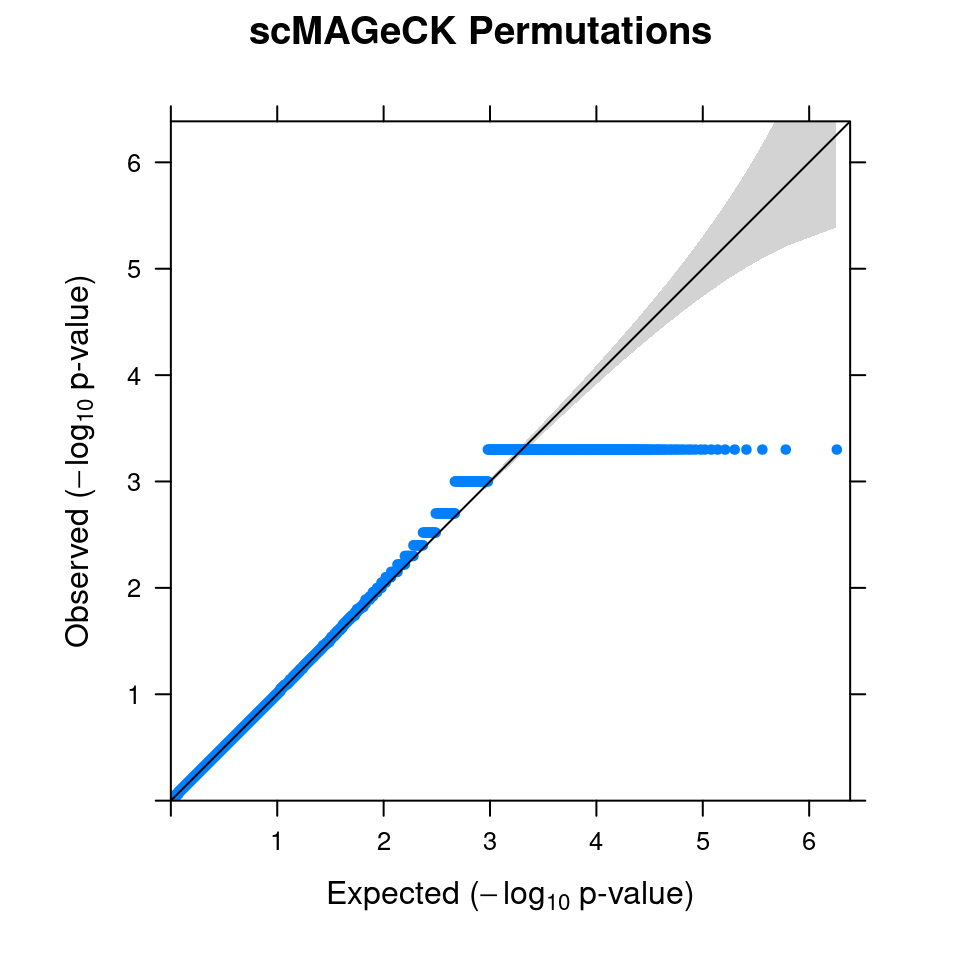
2.2 Original scMAGeCK result
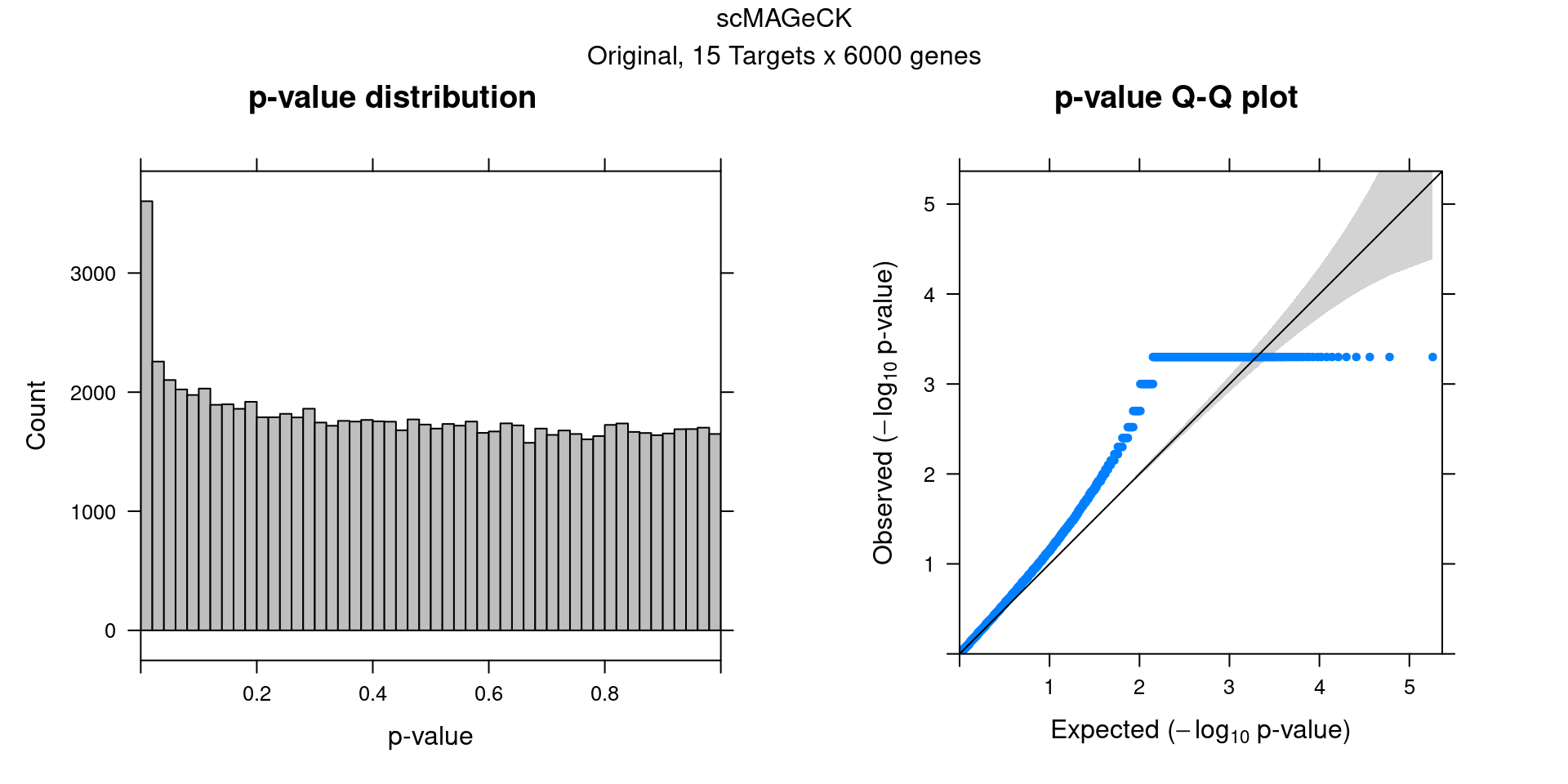
3 DESeq2 Permutation
Cell labels in the expression data were permuted randomly so that they are no longer correlated with the knock-down conditions. Then DESeq2 test was performed under each guide.
In total, 10 random permutation rounds like this were conducted.
3.1 Combined from 10 permutations
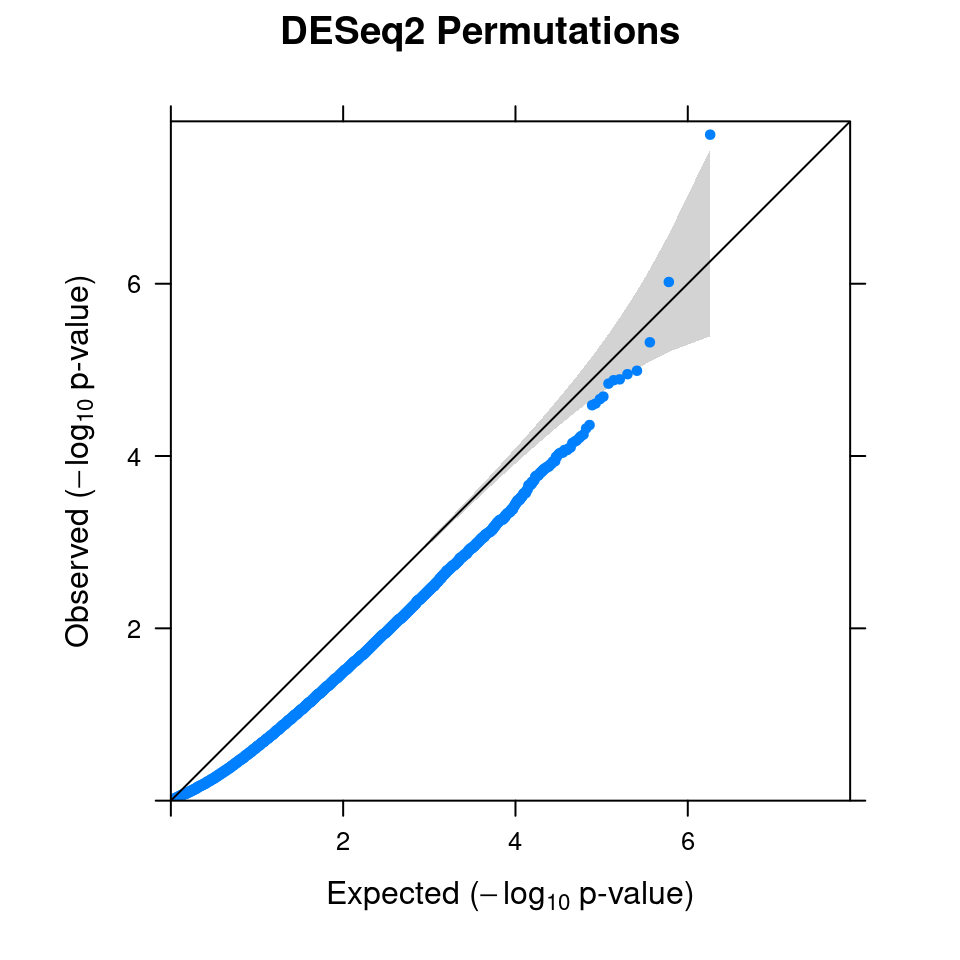
3.2 Original DESeq2 DGE result
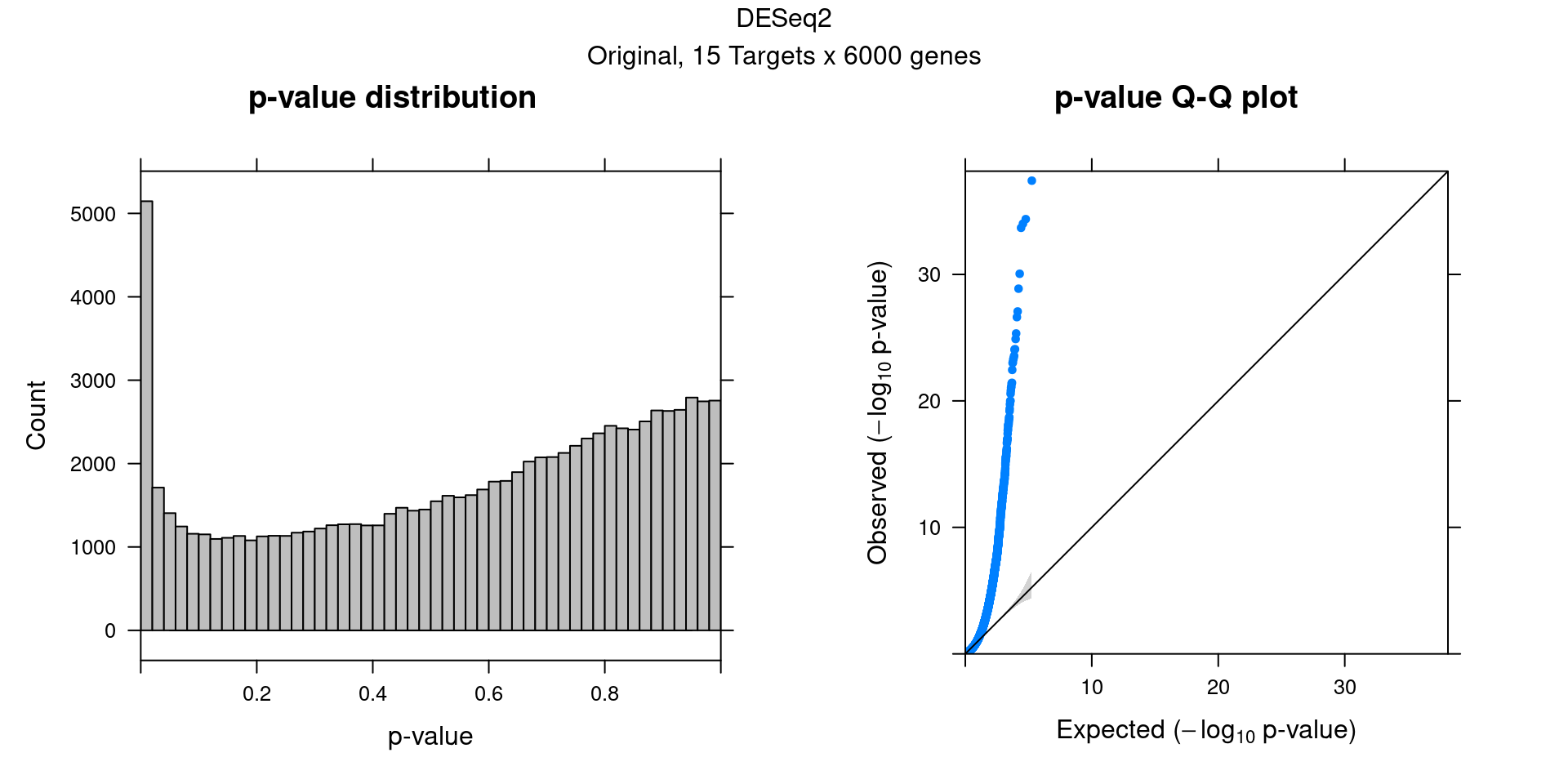
4 MAST Permutation
Cell labels in the expression data were permuted randomly so that they are no longer correlated with the knock-down conditions. Then MAST likelihood ratio test was performed under each guide.
In total, 10 random permutation rounds like this were conducted.
4.1 Combined from 10 permutations
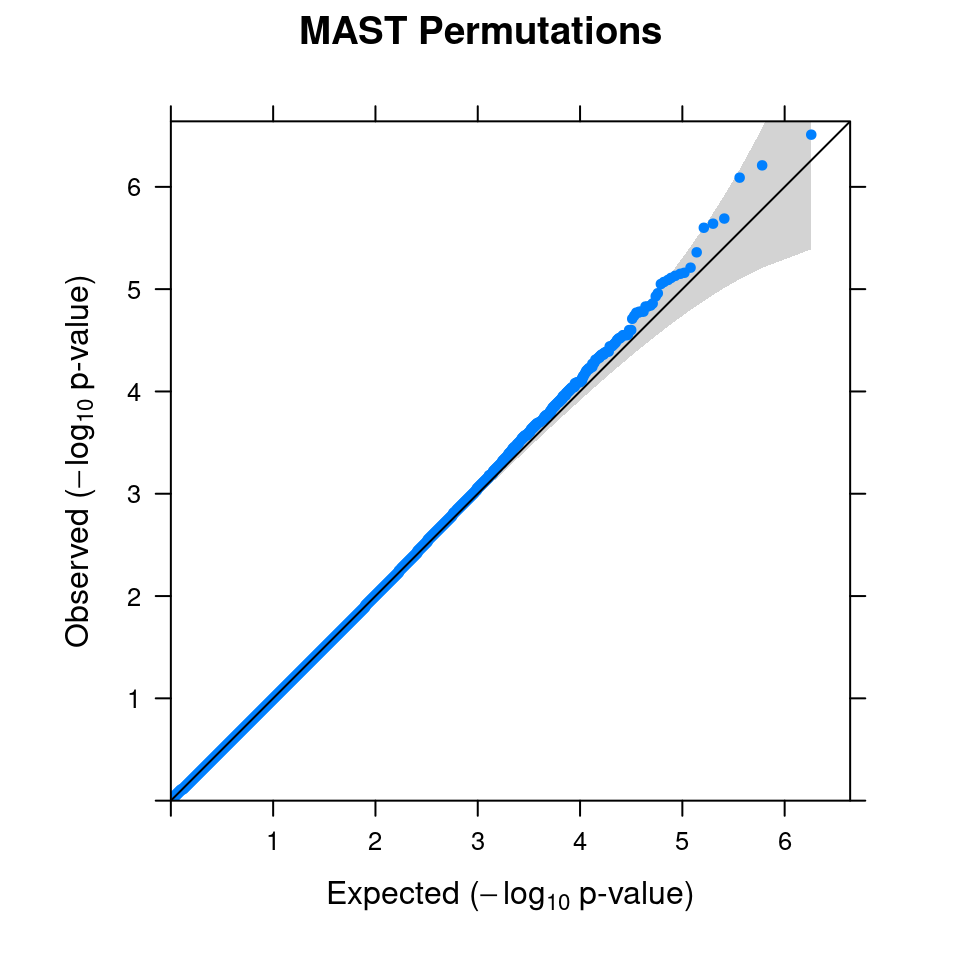
4.2 Original MAST DGE result
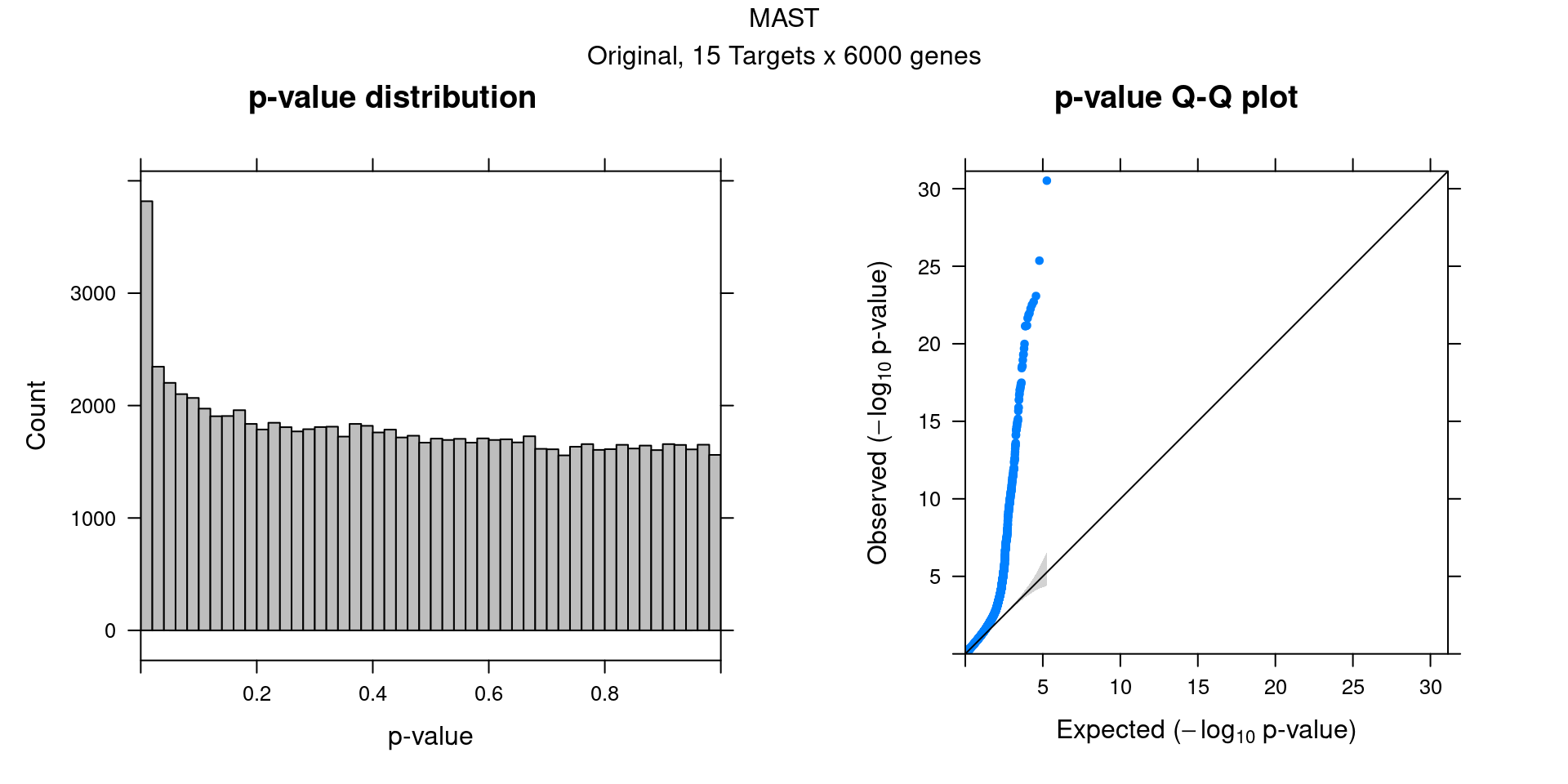
5 SCEPTRE
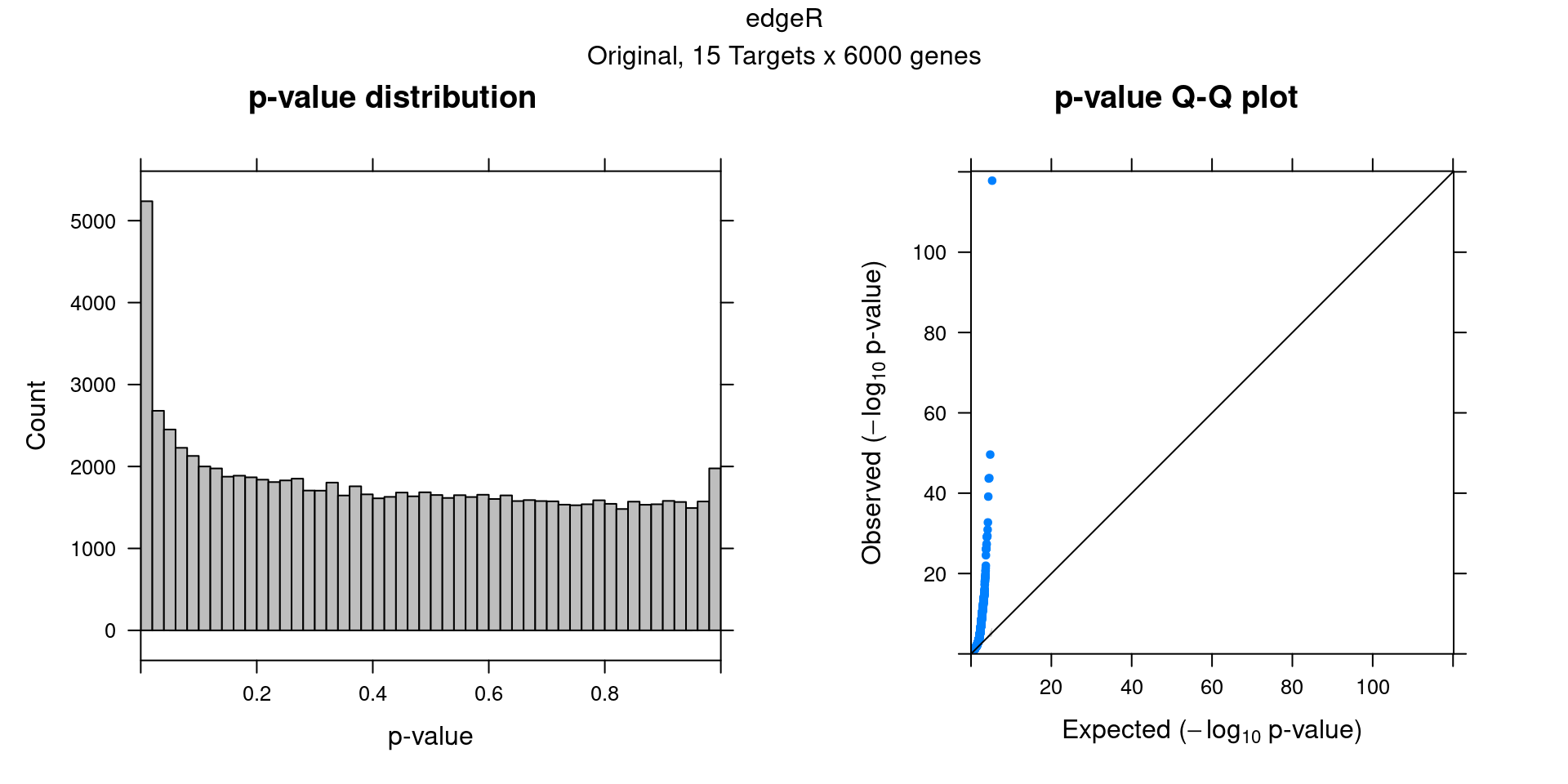
6 edgeR Permutation
Cell labels in the expression data were permuted randomly so that they are no longer correlated with the knock-down conditions. Then edgeR QLF test was performed under each guide.
In total, 10 random permutation rounds like this were conducted.
6.1 Combined from 10 permutations
6.2 Original edgeR DGE result